Professional Protein Identification Services
Professional Protein Identification Services
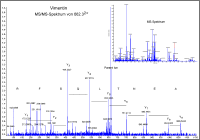
Protein identification by MSMS peptide fragmentation fingerprint (PFF) analysis
Proteome Factory offers protein identification by most modern mass spectrometric techniques. To ensure highest sensitivity and reliablilty Proteome Factory uses high resolution nanoLC-ESI-MS/MS which allows unequivocal identification of one and more proteins within a sample. This is very suitable for analysis of protein bands from SDS-PAGE, protein complexes and other low or medium complex protein samples.
Protein identification results will be provided electronically in one report including protein name, amino acid sequence and molecular weight.
We provide
The protein identification service is applied to protein samples delivered as protein gel spot, protein gel band, lyophilzed, in solution or bound to blotting membrane. Proteins are either analyzed by nanoLC-ESI-MSMS. The latter also identification of several protein within a sample due to peptide separation by nanoLC.
You receive
A result report of protein identification will be send by email or provided via SSL-server as a PDF file containing the results of protein database search.
How to order
Please, contact us and ship the sample together with our protein identification sample submission form signed by you. Below information are provided about sample requirements and shipping. Our sample submission form for protein identification by nanoLC-ESI-MSMS can be downloaded here
.
Express Services and Pricing
Common question in protein identifcation
1. MS compatible staining methods for protein gels
Protein gel staining methods which do not cause irreversible modification of proteins are compatible with protein identification by mass spectrometry like the following: Coomassie Brilliant Blue CBB-R250 and CBB-G250 and fluorescent staining like the ready-to-use FireRuth fluorescent staining solution. If you need a high sensitive silver staining we recommend the FireSilver protein gel staining kit which is more compatible to subsequent protein identification by MS than commonly used methods. Because conventional silver staining methods contain glutaraldehyde or high concentrations of formaldehyde which causes irreversible modification and crosslinking of proteins. These modified protein are badly to identify.
2. Keratin and other contaminations of protein samples
Keratin is a hair and skin protein which occurs almost everywhere in dust and it can easily contaminate lab equipment and reaction tubes because of electrostatic charging. Therefore we recommend using only RNase-free reaction tubes and pipetting tips. These tubes and lab dishes used for staining etc. should be well washed by 0.2 µm filtered Milli-Q water before use. Also it is highly recommendable to use a clean bench to prevent direct contamination of protein gels by dust and skin particles.
Recommendations for sample shipping
- Fill protein identification order form with information about the samples (e.g. sample name, sample type, MW, estimated protein amount, staining method) and the requested type of MS analysis, sign the form.
- Inform us by email or phone that your sample will be shipped.
- Transfer your protein spot or band in a 0.5 ml reaction tube without addition of any liquid.
- Seal your sample in a reaction tube by plastic wrap.
- Liquid samples should be shipped in a frozen state. Protein bands and protein spots can be shipped at room temperature.
- Ship the samples in a padded envelope or box together with the protein identification order form.
Shipping address
Proteome Factory AG
Magnusstr. 11
DE-12489 Berlin
Data security and secrecy
Proteome Factory takes security of your data and research results seriously. Therefore critical data are only internally used by our experts involved in the analysis task or research project. Communication via internet and email can be performed by using common safety and encryption techniques on request. Your data will be stored on redundant and fail-proof data storage servers.
On request a non-disclosure agreement can be signed.
Express Service
Express Service: Protein identification analysis by nanoLC-ESI-MSMS within days
We offer an Express Service for protein identification within 2-4 days after arrival of your sample.
Next-Day Service: Protein identification analysis by nanoLC-ESI-MSMS within a day.
We offer a Next-Day Service for protein identification within one day after arrival of your sample. The sample must arrive on a working day before 12am. We recommend for this service an overnight shipping service with guaranteed delivery next day before noon. This service is only available by contacting us in advance to ensure rapid sample processing.
Protein Identification - Prices for Europe*
(effective 1st October 2023)
Protein identification by nanoLC-ESI-MSMS
High sensitive protein identification by nanoLC-ESI-MSMS. This protein identification service is recommended for protein bands from PAGE gels, protein spots from 2DE gels and simple protein mixtures. nanoLC-ESI-MSMS protein identification results mostly in better protein identification and higher confidential sequence coverages than MALDI-MS analyses due to nanoLC separation of the proteolytical peptides. This yields much more MSMS peptide fragmentation fingerprint analyses resulting in mostly significantly higher protein identification scores by database searching. By default the MSMS data are searched against NCBInr protein database. Additional databases or databases provided by customer can be used upon request.
High resolution nanoLC-ESI-MSMS
Protein identification |
Price per sample [EUR] Standard Service (1-2 weeks) |
||
|
for samples with one to a few proteins (short gradient) |
Samples containing 1-10 proteins (short LCMS gradient, applies for analysis of Coomassie-stained PAGE bands or 2DE spots) | Medium complex samples or samples in solution (regular LCMS gradient) | Complex samples with many proteins or large dynamic range (long LCMS gradient) |
| 1 sample | 300 | 550 | 850 |
| 2 - 9 samples | 250 | 500 | 750 |
| 10 - 24 samples | 225 | 450 | 650 |
| 25 or more samples | inquire | inquire | inquire |
| Express Service (3-4 days) | +50% | ||
| Next-Day Service (1 day) | +100% |
Add-On services for protein identification
Further protein characterisation
LCMS-based proteomics, quantitative proteomics
| Price per sample [EURO] |
|
| iTRAQ based quantitative proteomics | on request |
| MudPIT proteomics | on request |
nanoLC-ESI-MSMS based proteomics |
on request |
Protein Profiling for 2DE gels
Protein Profiling of 2DE gels by nanoLC-ESI-MSMS
Protein profiling of protein spots from 2DE gels by nanoLC-ESI-MSMS ensures an high sensitive protein identification. The protein spots are cut automatically by using Proteome Factory’s HTS spot picking robot spotXpress according to customer’s spot list. Next the protein samples are processed by SpotDigest 96 with subsequent tryptic digest and nanoLC-ESI-MSMS analysis. The MSMS peptide fragmentation fingerprint (PFF) data are automatically searched against NCBInr protein database. Alternative database can be used on request. The results are transferred to the customer via email or SSL server.
This service requires a minimum of 50 protein spots to be analyzed.
|
at least 50 protein spots: 150 EURO* per spot at least 100 protein spots: 100 EURO* per spot |
*All price do not include VAT. In Germany (EU) 19% VAT has to be added.
** If we do not obtain any phosphopeptide or glycopeptide data, respectively, we will only charge you for the standard protein identification analysis by nanoLC-ESI-MSMS.
Discounts for multiple samples apply only for samples sent in together.